The Rio Piedras campus of the UPR published a short article on the IDI-BD2K project November 4th.
Author: humberto
Get ready for IDI-BD2K
We were awarded the IDI-BD2K grant, and are getting ready to select students for next year. If you want to participate, try to select the courses you need to get up to speed during pre-registration.
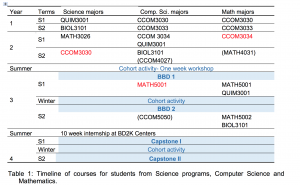
Here’s a copy of the table showing the courses we want you to have. If you have questions, speak to one of the participating researchers.
Roche-Lima, Abiel – Machine Learning to Predict Biological Networks.
Dr. Roche-Lima has been working on machine learning methods, based on kernels, to predict biological networks. He proposed a new framework, called Pairwise Rational Kernel (PRK), to manipulate sequence data represented as finite-state transducers (FSTs). By combining PRKs with supervised learning methods, biological network interactions have been predicted. As kernel methods are used, disparate type of data can be combined to find general relations. Using finite-state transducers, large amount of sequence data can be efficiently represented, processed and analyzed, improving the performance of the algorithms. Dr. Roche-Lima has been working and collaborating with bioinformatics studies at University of Manitoba, Canada, to predict biological interactions in several bacteria species. He is currently working at Medical Science Campus, University of Puerto Rico, where large volume of sequence data, from several projects, are being generated. Students in his lab will learn how to represent, manipulate and analyze these data using the existing frameworks and machine learning methods. As well, students will develop new computational tools using these techniques.
Due to his experience working with predicting models and biological sequence data, Dr. Roche-Lima brings to the project the ability to develop computational tools for analyzing and processing big sequence data. It can be used to predict biological network interactions, but also it can be extended to any other string data, such as text data in social network interactions.
Reunion informativa
Estimado estudiante,
Hoy como nunca antes la investigación biomédica está generando cantidades masivas de datos, cuyo análisis e interpretación tiene el potencial de producir dramáticos avances en nuestro conocimiento sobre la salud humana y sobre nuestra calidad de vida. El análisis de estos conjuntos masivos de datos (“Big Data”) require técnicas que combinan conocimientos en Biología, Química, Estadística, Ciencias de Cómputo y otras áreas.
Existe la posibilidad de que NIH apruebe una propuesta enviada por un grupo de profesores de la Facultad de Ciencias Naturales de la UPR-Rio Piedras para preparar estudiantes de diferentes concentraciones en investigación biomédica usando grandes cantidades de datos (“Big Data to Knowledge”- BD2K) Estos estudiantes tomarían una secuencia de cursos dependiendo de su concentración de origen, y también cursos enfocados en el manejo y análisis de “Biomedical Big Data”. Los mejores estudiantes de este grupo realizarán internados en laboratorios nacionales financiados por NIH.
Si eres estudiante de la Facultad de Ciencias Naturales y te interesa este reto:
te invitamos a una reunión informativa los días 10 y 12 de agosto de 2015 al medio dia en el anfiteatro A-211.
En esta reunion esperamos poder formar dos grupos de estudiantes. El primero con estudiantes comenzando su 2do, 3er o 4to año que estén avanzados en sus estudios y que puedan incorporarse al programa como un grupo piloto. El segundo, estudiantes de 1er a 3er año que puedan ir tomando los cursos necesarios para incorporarse al Programa el año entrante.
Pérez-Hernández, María-Eglée – Bayesian Biostatistics and its Applications in Life Sciences
Dr. Pérez Hernández is currently involved in the “Biostatistics, Epidemiology and Bioinformatics BEBiC Core” of the U54 Collaborative 5 year Grant between UPR and MDAnderson Cancer Center, where she is collaborating with Drs. Pericchi and Ortiz-Zuazaga. She is also collaborating with Dr. Acevedo in the development of Bayesian epidemiological models based on internet search information (Google Flu).
Dr. Pérez-Hernández has made contributions on Bayesian Statistics, especially on Bayesian Robustness and Objective Bayesian Methods. She has a long history of successful interdisciplinary work with researchers in biomedical sciences and ecology, including statistical support for development of rotavirus vaccines and statistical support for studies on Helicobacter pylori.
Pericchi, Luis – Bayesian Statistics in Cancer, Cardiovascular Disease and Health Econometrics
Dr. Pericchi has currently three long term projects that involve big data from Puerto Rico, and that require exploratory data analysis, modeling, inference and prediction. Currently he is the Co-PI of the “Biostatistics, Epidemiology and Bioinformatics BEBiC Core” of the U54 Collaborative 5 year Grant between the University of Puerto Rico and MDAnderson Cancer Center. He is collaborating with Drs. Perez-Hernandez and Ortiz-Zuazaga and directing students to search for predictive models of prostate cancer severity that involve over 800 patients and around 600 potential explanatory variables. Another aspect of his cancer-related research deals with the design of multidimensional engineering experiments for alternative cancer treatments to radio- and chemo- therapies that give rise to response surfaces in several dimensions. Regarding heart disease and stroke, he has worked with the School of Medicine Endowed Health Services Research Center, and a database of cardiovascular diseases in Puerto Rico was established with several possible explanatory variables, giving rise to several potential data science projects. Regarding health econometrics and related fields, Dr. Pericchi has been directing projects to capture masses of information of credit behavior in Puerto Rico, as well its modeling.
Dr. Pericchi has a long trajectory on different aspects of Bayesian Statistics, but especially in: Foundations of Decision Theory, Model Selection, Bayesian Robustness, Bayesian Treatment of Conflicting Evidence and Applications to Statistics of Extremes, Detection of Fraud, Medical Diagnoses and Clinical Trials. He is an elected member of ISBA: International Society for Bayesian Analysis and the current president of its Section of Objective Bayes.
Ortiz-Zuazaga, Humberto G. – Bioinformatics of Gene Expression
Dr. Ortiz-Zuazaga has developed novel methods of measuring gene expression from microarray and second-generation sequencing data, and determining regulatory gene networks from this data. He already has established successful collaborations with scientists in biomedical research using Big Data, in this award, he will continue to grow these research collaborations, bringing his quantitative and algorithmic skills to bear on novel biomedical problems. Due to his experience in multiple fields, Dr. Ortiz-Zuazaga is uniquely qualified to abstract the basic algorithmic challenges in many biological problems, and can help translate biological questions into data analysis algorithms. Students in his lab will adapt probabilistic data structures to the task of detecting differential gene expression in de-novo RNA-seq experiments, and use these and other data sets to model gene regulatory networks using bioinformatic and statistical methods.
Dr. Ortiz-Zuazaga brings to the project extensive experience in computational biology, ranging from data analysis to modelling and simulation and visualization.
Ordóñez, Patricia – Visualization, Machine Learning, and Biomedical Informatics Education
Professor Patricia Ordóñez has been developing a real-time visualization for Intensive Care Unit Data for over 7 years. She will be working with the MIT Laboratory of Computational Physiology in a summer sabbatical in 2015 to incorporate her visualization into their soon-to-be publically available database of streaming physiological data. As part of this grant, she envisions working with Dr. Harry Hochheiser at the University of Pittsburgh on the development and assessment of this project. She would like to incorporate his research on time boxes for univariate time series into multivariate time series of vital sign data. He would serve as a mentor in this project to improve the user experience.
Patricia Ordóñez is the founder of the Symposium of Health Informatics in Latin America and the Caribbean (SHILAC) that began in 2013 with an emphasis on defining common health care problems in LAC and finding innovative informatics solutions. The second SHILAC accompanied by the first Hacking Medicine in the Caribbean will occur in November 2015 in San Juan. Her contacts in Latin America and the Caribbean with leaders in biomedical informatics will serve as mentors for faculty at UPR-RP. Her expertise in working with visualization and machine learning in multivariate times series to develop clinical decision systems make her an ideal candidate for the program since she is attempting to incorporate her research into streaming databases.
Massey, Steve E. – Meta-metabolomic Network Analysis of Metagenomic Data from Diverse Habitats from Around the World
Dr Massey has been developing methods to assess metabolic flux through a microbial community from shotgun metagenomic data, by reconstructing ‘meta-metabolomic networks’ which show the relative abundance of genes encoding enzymes involved in the different metabolic pathways present. The approach involves large scale Blast searching of millions of individual sequences using grid computing, assignment of metabolic function to the identified sequence homologs, calculation of relative redundancy from the dataset, and calculation of overall flux using the kinetic rate constants of reference enzymes taken from the literature. The overall aim of this project is to assess differences in carbon flux from diverse habitats around the world, with an emphasis on methanogenesis. Data will be obtained from the MG-RAST database and selected for variation in latitude, temperature, and aerobicity. Students will learn a range of command line driven techniques for conducting both local and remote analyses, and will learn how to manage and parse very large data sets.
Dr. Massey is a bioinformatician with a wide range of interests, in genome evolution, metagenomics, organismal complexity, genetic code evolution, evolutionary medicine and ancient DNA.
Koutis, Yiannis – Algorithm Development for Image Segmentation
Dr. Koutis and his former MS student Richard Garcia-Lebron have developed new optimization-based methods for semi-automatic segmentation of neurons in EM images. These methods produce segmentations whose quality comes close to that of human experts. These methods require very little human intervention, and complete the segmentation in a small fraction of the time needed for manual segmentation. At the heart of these algorithms are recently discovered solvers and optimization techniques in which Dr Koutis has been a key contributor. This ongoing project offers many opportunities for undergraduate students with different sets of skills and interests and at various levels. Conversely, the contribution of undergraduate students is beneficial for the project as it can provide the lower-level support for more advanced students, and a stream of potential contributors to the larger field of Connectomics, under the auspices of NIH’s BRAIN initiative.
Dr. Koutis bring to the project knowledge in theoretical computer science, with expertise in spectral graph theory, numerical linear algebra and parameterized algorithms for hard combinatorial problems.